|
11/1/2022 0 Comments Tutorial de proteus 8 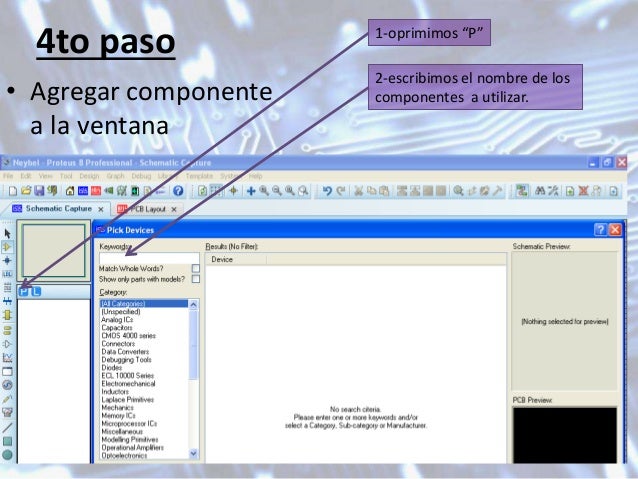 The choice of column to aggregate protein data is controlled by parameter l in the makePeptideTable function. We can change this default approach and, instead of razor proteins, we can use protein groups, as listed in Proteins column of the evidence file (this column is renamed to protein_group by readEvidenceFile to avoid confusion). This means that for a given protein makeProteinTable would aggregate data from all peptides having this protein ID in the Leading razor protein column. Here, we added a replicate column, but other information, in particular describing batch effects, can be very useful.īy default protein intensities are aggregated from peptide intensities based on the Leading razor protein column from the evidence file (this column is renamed to protein by readEvidenceFile for simplicity). Other columns can be added and used for the downstream analysis. In our example there are two conditions, hence two values in this column. This information will be used in differential expression. Sample identifies a given experiment/measure uniquely, so sample names must be unique.Ĭolumn condition contains condition names. For clarity, we recommend a short name consisting of a condition and replicate (in a simple design where such decomposition is possible). In multiplexed data (e.g. TMT) there can be several measurements columns.Ĭolumn sample should contain (short) names corresponding to a given experiment and measure. In case of the label-free experiment there is only one measure column, “Intensity”. # "AIF MS/MS IDs" "Deamidation (NQ) site IDs" # "Number of scans" "Number of isotopic peaks" # "Retention time calibration" "Match time difference" # "Calibrated retention time start" "Calibrated retention time finish" # "Retention length" "Calibrated retention time" # "Uncalibrated mass error " "Uncalibrated mass error " # "Uncalibrated - Calibrated m/z " "Uncalibrated - Calibrated m/z " # "Leading proteins" "Leading razor protein" The data are distributed in a package proteusLabelFree, which needs installing and loading first: We use an example data set from an unpublished experiment by Katharina Trunk, Sarah Coulthurst, Julien Peltier and Matthias Trost. TUTORIAL DE PROTEUS 8 HOW TOThis tutorial demonstrates how to analyse data from label free MS/MS experiment in Proteus. 
6.1 Proteins present in only one condition.5.5 Reading MaxQuant’s protein groups file.If you do not allow these cookies, you will experience reduced relevant content. 
They do not store directly personal information, but are based on uniquely identifying your browser and internet device. They may be used by Analog Devices to build a profile of your interests and show you relevant content on our site. Targeting Cookies: These cookies may be set through our site by Analog Devices and our service providers. If you do not allow these cookies we will not know when you have visited our site, and will not be able to monitor its performance. All information these cookies collect is aggregated and therefore anonymous. They help us to know which pages are the most and least popular and see how visitors move around the site. Performance Cookies: These cookies allow us to count visits and traffic sources so we can measure and improve the performance of our site. If you do not allow these cookies then some or all of these services may not function properly. They may be set by us or by third party providers whose services we have added to our pages. Functional Cookies: These cookies enable the website to provide enhanced functionality and personalization. These cookies do not store any personally identifiable information. You can set your browser to block or alert you about these cookies, but some parts of the site will not then work. They are usually only set in response to actions made by you which amount to a request for services, such as setting your privacy preferences, logging in or filling in forms. Strictly Necessary Cookies: (Always Active) These cookies are necessary for the website to function and cannot be switched off in our systems. After we finish updating our website, you will be able to set your cookie preferences. Analog Devices is in the process of updating our website. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
